Experiment 1 - Encoding precision
Contents
import pathlib
from typing import List
import gym
import gym_multiplexer # noqa: F401
import lcs.agents.racs as racs
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import pandas as pd
from IPython.display import display, HTML
from lcs.representations.RealValueEncoder import RealValueEncoder
from tabulate import tabulate
from myst_nb import glue
from src.bayes_estimation import bayes_estimate
from src.commons import NUM_EXPERIMENTS
from src.decorators import repeat, get_from_cache_or_run
from src.utils import build_plots_dir_path, build_cache_dir_path
from src.visualization import PLOT_DPI
plt.ioff() # turn off interactive plotting
root_dir = pathlib.Path().cwd().parent.parent.parent.parent
cwd_dir = pathlib.Path().cwd()
plot_dir = build_plots_dir_path(root_dir) / cwd_dir.parent.name / cwd_dir.name
cache_dir = build_cache_dir_path(root_dir) / cwd_dir.parent.name / cwd_dir.name
TRIALS = 20_000
USE_RAY = True
def encode(p, bits):
return int(RealValueEncoder(bits).encode(p))
def metrics_to_df(metrics: List) -> pd.DataFrame:
lst = [[d['trial'], d['reward'], d['population'], d['reliable']] for d in metrics]
df = pd.DataFrame(lst, columns=['trial', 'reward', 'population', 'reliable'])
df = df.set_index('trial')
df['phase'] = df.index.map(lambda t: "explore" if t % 2 == 0 else "exploit")
return df
def average_experiment_runs(runs_dfs: List[pd.DataFrame]) -> pd.DataFrame:
return pd.concat(runs_dfs).groupby(['trial', 'phase']).mean().reset_index(level='phase')
def single_experiment(env_provider, encoder_bits, trials):
env = env_provider()
env.reset()
def _metrics(agent, environment):
population = agent.population
return {
'population': len(population),
'numerosity': sum(cl.num for cl in population),
'reliable': len([cl for cl in population if cl.is_reliable()])
}
cfg = racs.Configuration(
classifier_length=env.observation_space.shape[0],
number_of_possible_actions=env.action_space.n,
encoder=RealValueEncoder(encoder_bits),
metrics_trial_frequency=5,
user_metrics_collector_fcn=_metrics,
epsilon=1.0, # no biased exploration
do_ga=True,
theta_r=0.9,
theta_i=0.2,
theta_ga=100,
cover_noise=0,
mutation_noise=0.25,
chi=1.0,
mu=0.1)
# create agent
agent = racs.RACS(cfg)
# run computations
metrics = agent.explore_exploit(env, trials)
return metrics_to_df(metrics)
def plot(df, fig_title, plot_filename=None):
major_ticker_freq = 4000
# separate explore/exploit data
explore_df = df[df['phase'] == 'explore']
exploit_df = df[df['phase'] == 'exploit']
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 5))
fig.suptitle(fig_title, fontsize=14)
# plot 1 - average reward
explore_df['reward'].rolling(window=50).mean().plot(label='explore', ax=ax1)
exploit_df['reward'].rolling(window=50).mean().plot(label='exploit', ax=ax1)
ax1.axhline(y=500, color='black', linestyle='--', linewidth=1)
ax1.axhline(y=1000, color='black', linestyle='--', linewidth=1)
ax1.set_title('Average reward')
ax1.set_xlabel('Trial')
ax1.set_ylabel('Average reward')
ax1.set_ylim(400, 1100)
ax1.xaxis.set_major_locator(ticker.MultipleLocator(major_ticker_freq))
ax1.legend()
# plot 2 - number of classifiers
df['population'].plot(label='population', ax=ax2)
df['reliable'].plot(label='reliable', ax=ax2)
ax2.set_title("Classifiers evolution")
ax2.set_xlabel('Trial')
ax2.set_ylabel('# Classifiers')
ax2.xaxis.set_major_locator(ticker.MultipleLocator(major_ticker_freq))
ax2.legend()
if plot_filename:
fig.savefig(plot_filename, dpi=PLOT_DPI)
return fig
glue('32_e1_trials', TRIALS, display=False)
Experiment 1 - Encoding precision¶
The impact of setting the encoder bits value when forming interval predicates was validated on the single-step rMPX environment. Due to potential computational complexity issues, the problem was scaled down to 3bit rMPX, where the perception vector is represented by four attributes (the first three bits are both the address and register, and the last one indicates whether the given answer was correct).
The impact of four different encoder bits values was examined by collecting metrics, including representative metrics of the average reward in each execution phase and the size of the evolved population.
In each trial of the experiment, the agent alternates between explore and exploit phases for the total of 20000 trials, which allows a discerning shift in performance over time. Moreover, to present coherent results and draw statistical inferences, each experiment is repeated 50 times and those independent runs are averaged.
Results¶
rACS parameters
\(\beta=0.05\), \(\gamma = 0.95\), \(\theta_r = 0.9\), \(\theta_i=0.2\), \(\epsilon = 1.0\), \(\theta_{GA} = 100\), \(m_u=0.1\), \(\chi=1.0\), \(\epsilon_{cover} = 0\), \(\epsilon_{mutation}=0.25\).
def rmpx3bit_env_provider():
import gym_multiplexer # noqa: F401
return gym.make('real-multiplexer-3bit-v0')
@get_from_cache_or_run(cache_path=f'{cache_dir}/rmpx_3bit/encoding_1bit.dill')
@repeat(num_times=NUM_EXPERIMENTS, use_ray=USE_RAY)
def run_3bit_rmpx_1bit_encoding():
return single_experiment(rmpx3bit_env_provider, encoder_bits=1, trials=TRIALS)
@get_from_cache_or_run(cache_path=f'{cache_dir}/rmpx_3bit/encoding_2bit.dill')
@repeat(num_times=NUM_EXPERIMENTS, use_ray=USE_RAY)
def run_3bit_rmpx_2bit_encoding():
return single_experiment(rmpx3bit_env_provider, encoder_bits=2, trials=TRIALS)
@get_from_cache_or_run(cache_path=f'{cache_dir}/rmpx_3bit/encoding_3bit.dill')
@repeat(num_times=NUM_EXPERIMENTS, use_ray=USE_RAY)
def run_3bit_rmpx_3bit_encoding():
return single_experiment(rmpx3bit_env_provider, encoder_bits=3, trials=TRIALS)
@get_from_cache_or_run(cache_path=f'{cache_dir}/rmpx_3bit/encoding_4bit.dill')
@repeat(num_times=NUM_EXPERIMENTS, use_ray=USE_RAY)
def run_3bit_rmpx_4bit_encoding():
return single_experiment(rmpx3bit_env_provider, encoder_bits=4, trials=TRIALS)
# run computations
rmpx3bit_encoding_1bit_results = run_3bit_rmpx_1bit_encoding()
rmpx3bit_encoding_2bit_results = run_3bit_rmpx_2bit_encoding()
rmpx3bit_encoding_3bit_results = run_3bit_rmpx_3bit_encoding()
rmpx3bit_encoding_4bit_results = run_3bit_rmpx_4bit_encoding()
glue("f1", plot(average_experiment_runs(rmpx3bit_encoding_1bit_results), '3bit rMPX with 1bit UBR encoding', plot_filename=f'{plot_dir}/rmpx_3bit_encoding_1bit.png'), display=False)
glue("f2", plot(average_experiment_runs(rmpx3bit_encoding_2bit_results), '3bit rMPX with 2bit UBR encoding', plot_filename=f'{plot_dir}/rmpx_3bit_encoding_2bit.png'), display=False)
glue("f3", plot(average_experiment_runs(rmpx3bit_encoding_3bit_results), '3bit rMPX with 3bit UBR encoding', plot_filename=f'{plot_dir}/rmpx_3bit_encoding_3bit.png'), display=False)
glue("f4", plot(average_experiment_runs(rmpx3bit_encoding_4bit_results), '3bit rMPX with 4bit UBR encoding', plot_filename=f'{plot_dir}/rmpx_3bit_encoding_4bit.png'), display=False)
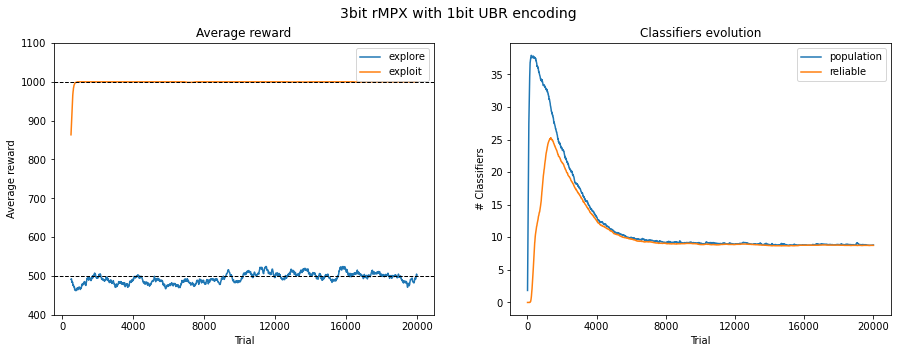
Fig. 3.2 Performance in 3bit rMPX UBR with 1bit. The reward is averaged across 50 last runs.¶
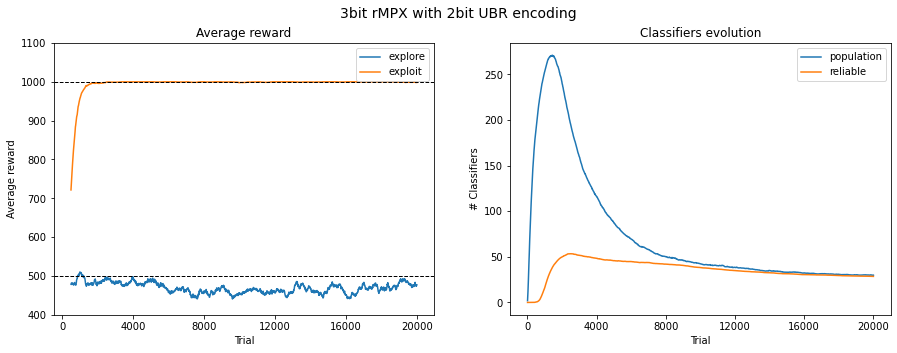
Fig. 3.3 Performance in 3bit rMPX UBR with 2bit. The reward is averaged across 50 last runs.¶
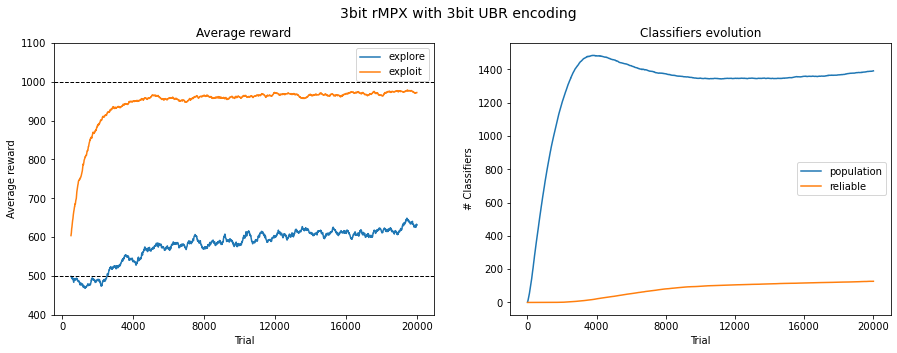
Fig. 3.4 Performance in 3bit rMPX UBR with 3bit. The reward is averaged across 50 last runs.¶
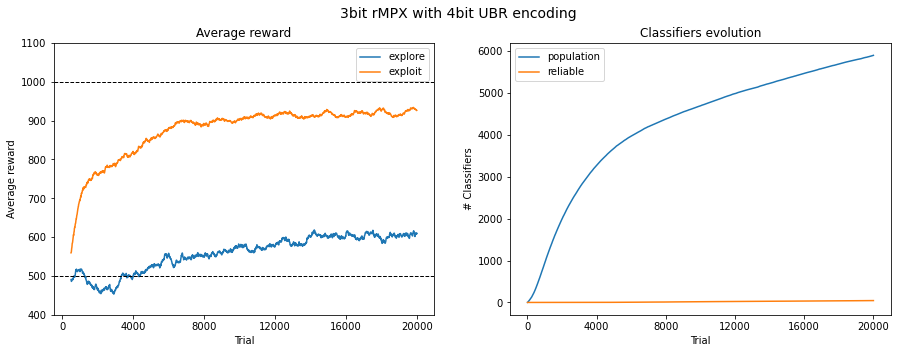
Fig. 3.5 Performance in 3bit rMPX UBR with 4bit. The reward is averaged across 50 last runs.¶
Statistical verification¶
To statistically assess the population size, the posterior data distribution was modelled using 50 metric values collected in the last trial and then sampled with 100,000 draws. The average value from the last 100 exploit trials is considered a representative state of algorithm performance for the obtained reward.
@get_from_cache_or_run(cache_path=f'{cache_dir}/rmpx_3bit/bayes/population.dill')
def build_population_models(dfs: List[pd.DataFrame]):
query_condition = f'trial == {TRIALS}' # last phase
results = []
for df in dfs:
data_arr = df.query(query_condition)['population'].to_numpy()
bayes_model = bayes_estimate(data_arr)
results.append((bayes_model['mu'], bayes_model['std']))
return results
@get_from_cache_or_run(cache_path=f'{cache_dir}/rmpx_3bit/bayes/reliable.dill')
def build_reliable_models(dfs: List[pd.DataFrame]):
query_condition = f'trial == {TRIALS}' # last phase
results = []
for df in dfs:
data_arr = df.query(query_condition)['reliable'].to_numpy()
bayes_model = bayes_estimate(data_arr)
results.append((bayes_model['mu'], bayes_model['std']))
return results
def get_average_reward(dfs: List[pd.DataFrame], last_n_runs: int = 100):
results = []
for df in dfs:
avg_reward = df.query('phase == "exploit"').groupby('trial').mean().iloc[-last_n_runs:]['reward'].mean()
results.append(avg_reward)
return results
bayes_results_dfs = [
pd.concat(rmpx3bit_encoding_1bit_results),
pd.concat(rmpx3bit_encoding_2bit_results),
pd.concat(rmpx3bit_encoding_3bit_results),
pd.concat(rmpx3bit_encoding_4bit_results)
]
population_models = build_population_models(bayes_results_dfs)
reliable_models = build_reliable_models(bayes_results_dfs)
avg_rewards = get_average_reward(bayes_results_dfs)
bayes_table_data = [
['population of classifiers'] + [f'{round(r[0].mean(), 2)} ± {round(r[0].std(), 2)}' for r in population_models],
['reliable classifiers'] + [f'{round(r[0].mean(), 2)} ± {round(r[0].std(), 2)}' for r in reliable_models],
['reward from last 100 exploit runs'] + [f'{round(r, 2)}' for r in avg_rewards],
]
table = tabulate(bayes_table_data, headers=['', '1 bit', '2 bit', '3 bit', '4 bit'], tablefmt="html", stralign='right')
display(HTML(table))
| 1 bit | 2 bit | 3 bit | 4 bit | |
|---|---|---|---|---|
| population of classifiers | 8.8 ± 0.13 | 29.58 ± 0.71 | 1392.25 ± 25.68 | 5892.82 ± 86.8 |
| reliable classifiers | 8.75 ± 0.12 | 28.47 ± 0.54 | 127.08 ± 2.33 | 44.35 ± 1.6 |
| reward from last 100 exploit runs | 1000.0 | 998.6 | 974.6 | 927.6 |
Observations¶
The rACS algorithm implementation proved to be capable of solving 3bit rMPX problem with the hyper-plane decision surface. The performance of four distinct values of encoding resolutions are depicted in Figures 3.2, 3.3, 3.4, 3.5.
For this particular environment, the threshold mapping each real-value attribute into binary representation was set to \(0.5\); therefore, satisfying performance was obtained with very rough encoding values. The agent consistently exploited the environment with a single encoding bit by choosing the correct answer and converging the population size.
Raising the encoding bits should lead to similar results but slower. Due to the expansion of possible interval combinations, the learning component needs more time to evolve proper rules. This progression is observed in Figure 3.3 where 2 encoding bits are used. Similar performance is observed, but the final population size is almost three times larger due to the lack of rule compaction ability.
The problems are emphasized when using 3 and 4 encoding bits. The average reward in the exploit phase is still significantly greater than in explore one, meaning that the agent can pick up correct decisions most of the time, but the convergence of population size is becoming elusive. The algorithm is struggling with increasing population size, which, due to its nature, is a significant computational bottleneck.
Software packages used
import session_info
session_info.show()
Click to view session information
----- gym 0.21.0 gym_multiplexer NA lcs NA matplotlib 3.5.1 myst_nb 0.13.1 pandas 1.4.0 session_info 1.0.0 src (embedded book's utils module) tabulate 0.8.9 -----
Click to view modules imported as dependencies
PIL 8.4.0 arviz 0.11.2 asttokens NA attr 21.4.0 babel 2.9.1 backcall 0.2.0 beta_ufunc NA binom_ufunc NA bitstring 3.1.7 brotli NA cachetools 5.0.0 certifi 2021.10.08 cffi 1.15.0 cftime 1.5.2 charset_normalizer 2.0.10 click 7.1.2 cloudpickle 2.0.0 colorama 0.4.4 colorful 0.5.4 colorful_orig 0.5.4 cryptography 36.0.1 cycler 0.10.0 cython_runtime NA databricks_cli NA dataslots NA dateutil 2.8.2 debugpy 1.5.1 decorator 5.1.1 defusedxml 0.7.1 dill 0.3.4 docutils 0.16 entrypoints 0.3 executing 0.8.2 fastprogress 0.2.7 filelock 3.4.2 google NA greenlet 1.1.2 grpc 1.43.0 hiredis 2.0.0 idna 3.3 imagesize NA importlib_metadata NA ipykernel 6.7.0 ipython_genutils 0.2.0 ipywidgets 7.6.5 jedi 0.18.1 jinja2 3.0.3 jsonschema 3.2.0 jupyter_cache 0.4.3 jupyter_sphinx 0.3.2 jupyterlab_pygments 0.1.2 kiwisolver 1.3.2 linkify_it 1.0.3 markdown_it 1.1.0 markupsafe 2.0.1 matplotlib_inline NA mdit_py_plugins 0.2.8 mistune 0.8.4 mlflow 1.23.1 mpl_toolkits NA msgpack 1.0.3 myst_parser 0.15.2 nbclient 0.5.10 nbconvert 6.4.1 nbformat 5.1.3 nbinom_ufunc NA netCDF4 1.5.8 numpy 1.22.1 packaging 21.3 pandocfilters NA parso 0.8.3 pexpect 4.8.0 pickleshare 0.7.5 pkg_resources NA prompt_toolkit 3.0.26 psutil 5.9.0 ptyprocess 0.7.0 pure_eval 0.2.2 pvectorc NA pydev_ipython NA pydevconsole NA pydevd 2.6.0 pydevd_concurrency_analyser NA pydevd_file_utils NA pydevd_plugins NA pydevd_tracing NA pygments 2.11.2 pylab NA pymc3 3.11.4 pyparsing 3.0.7 pyrsistent NA pytz 2021.3 ray 1.9.2 redis 4.1.2 requests 2.27.1 scipy 1.7.3 semver 2.13.0 setproctitle 1.2.2 setuptools 60.5.0 six 1.16.0 socks 1.7.1 sphinx 4.4.0 sphinxcontrib NA sqlalchemy 1.4.31 stack_data 0.1.4 testpath 0.5.0 theano 1.1.2 tornado 6.1 tqdm 4.62.3 traitlets 5.1.1 typing_extensions NA uc_micro 1.0.1 unicodedata2 NA urllib3 1.26.8 wcwidth 0.2.5 xarray 0.21.0 yaml 6.0 zipp NA zmq 22.3.0
----- IPython 8.0.1 jupyter_client 7.1.2 jupyter_core 4.9.1 notebook 6.4.8 ----- Python 3.9.10 | packaged by conda-forge | (main, Feb 1 2022, 21:24:11) [GCC 9.4.0] Linux-5.13.0-30-generic-x86_64-with-glibc2.31 ----- Session information updated at 2022-02-24 12:35
